- 1革新铁路安全管理,RFID电子锁技术提升效率与防护
- 2adb 连接 显示 List of devices attached_list of devices attached eior9tlnhqtgpzyx device 1
- 3pycharm 常用插件,常用插件推荐_pycharm 插件
- 4NLP笔记(1)——深度学习和神经网络_nlp神经网络
- 5canal 同步数据_canal 数据同步
- 6苹果被大模型打得措手不及
- 7啊哈c语言读后感500字,《麦田里的守望者》读后感读书笔记500字五篇
- 8机器学习中的欠拟合和过拟合
- 9NLP之NER:商品标题属性识别探索与实践
- 10推荐十款开源测试开发工具(自动化、性能、造数据、流量复制)_开源测试工具
Chapter4 : Application of Artificial Intelligence and Machine Learning in Drug Discovery_applications of artifificial intelligence and mach
赞
踩
reading notes of《Artificial Intelligence in Drug Design》
1.Introduction
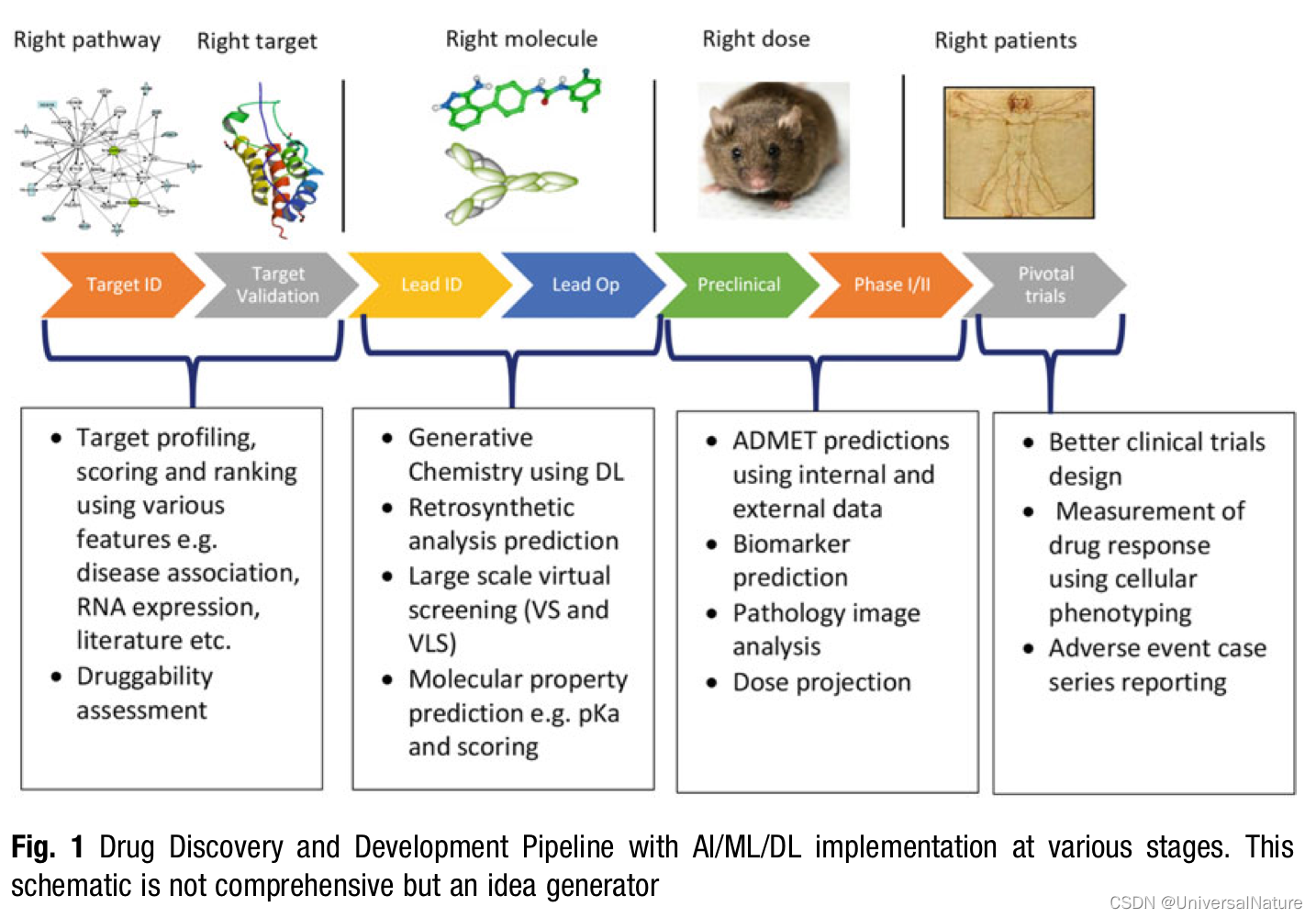
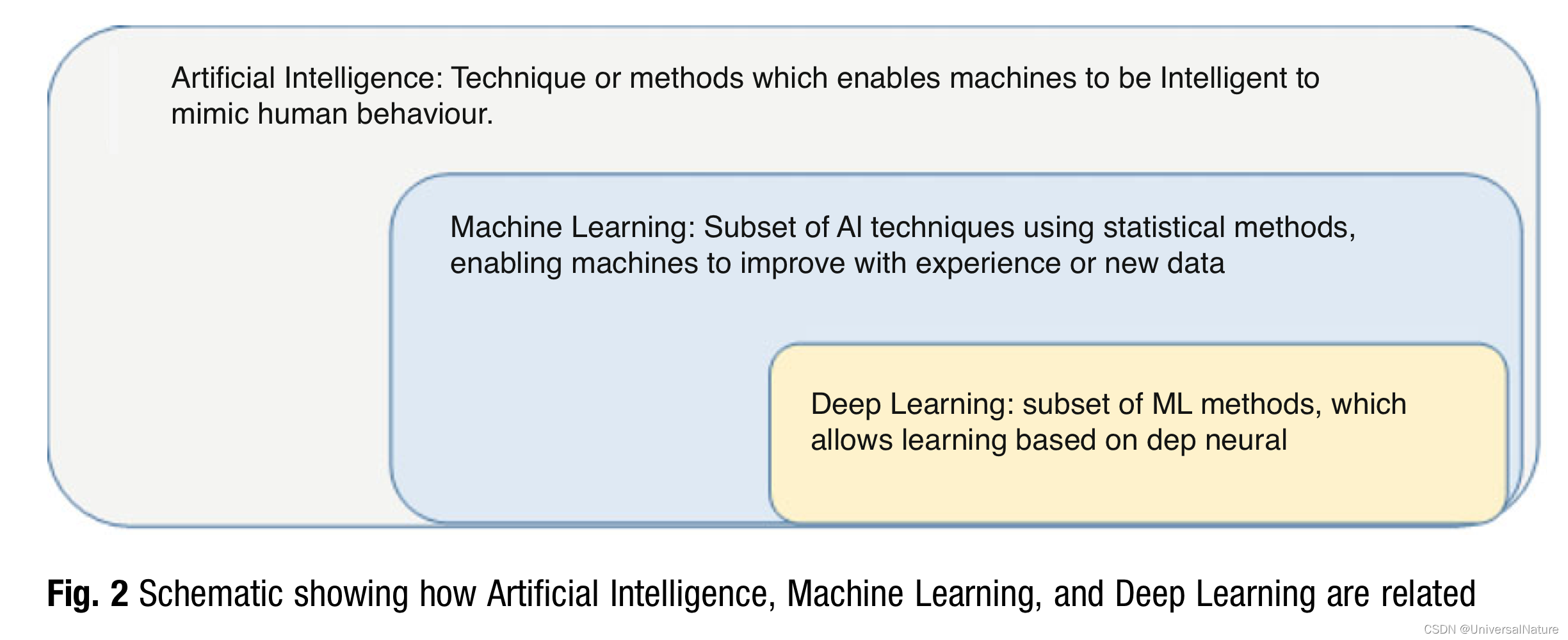
-
In addition to FAIR principles, Schneider et al. provide an excellent discussion on how data should also follow the ALCOA (Attributable, Legible, Contemporaneous, Original and Accurate) guidelines as defined by US FDA.
-
As a general principal when an opportunity or challenge is recognized within the drug discovery pipeline, we first ask ourselves if applying machine learning would be a good idea. Are there other methods that may be better as well as quicker to get us the desired information? This leads to investigating the actual use case as well as evaluating the amount and quality of data available for such application.
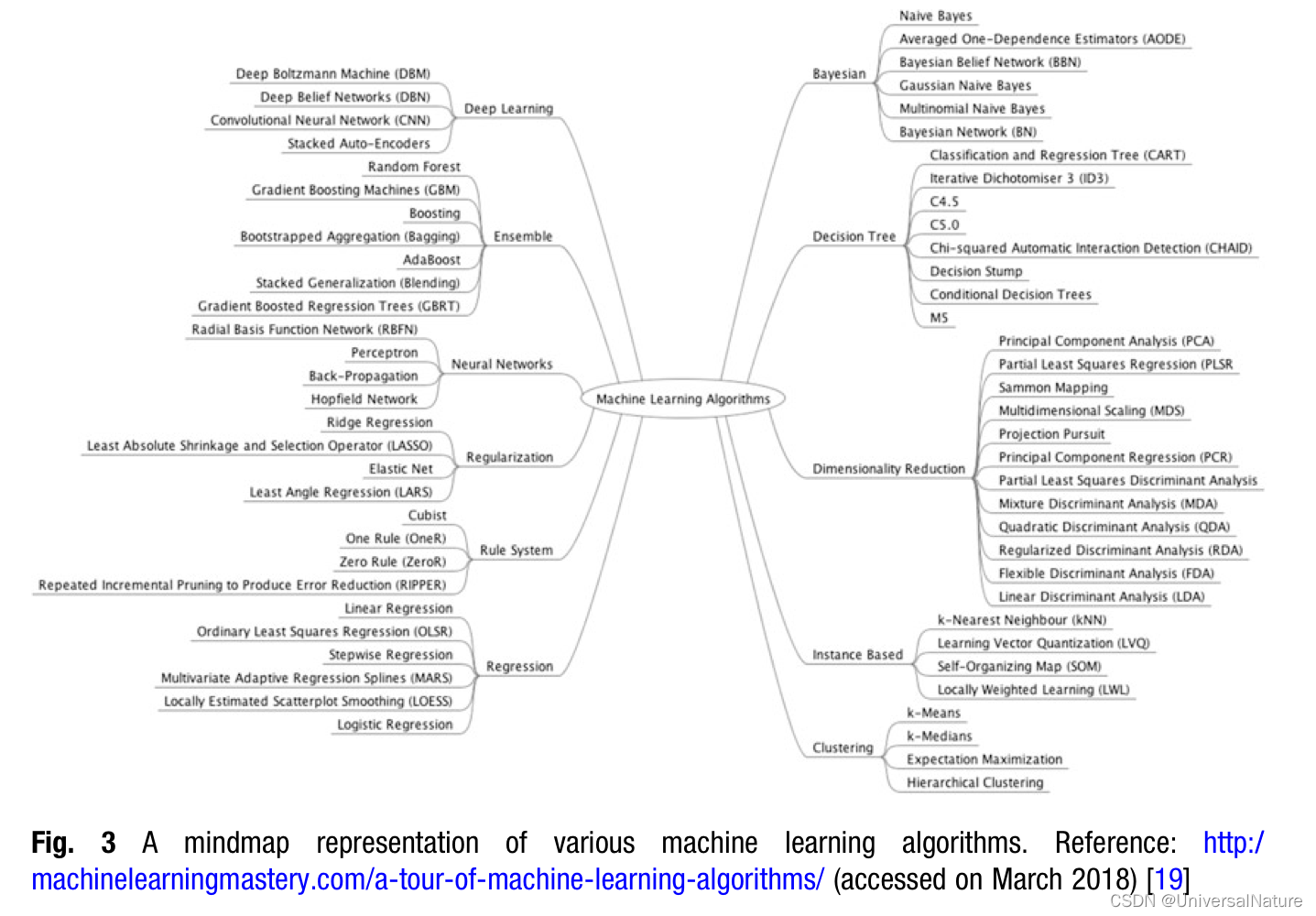
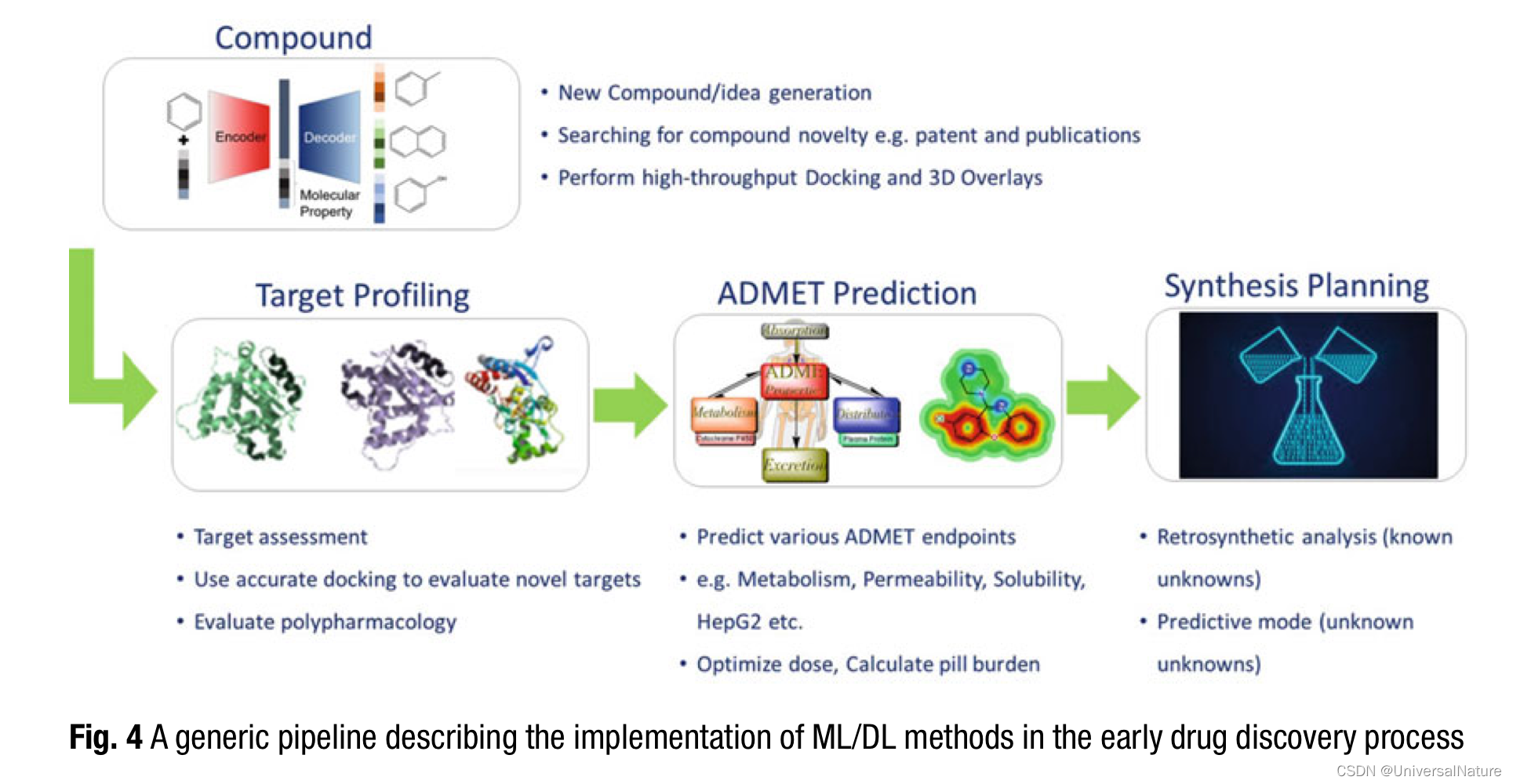
2.Generative Chemistry
-
Generative chemistry methods can combine scoring based on multiparameters to allow picking compounds that check most of the criteria as set by the project teams.
-
There has been work done to bring chemistry and biology close to each other by utilizing gene expression information in de-novo compound generation.
-
Potentially possible, it would be useful to allow retrosynthesis be part of the latent space during the generative chemistry process so that users can get synthetically viable compounds.
3.Target Profiling
- The next challenge at hand is target profiling or target assessment. This also includes predicting polypharmacology as well as off-target effects (including toxicity predictions).
- A wishful thinking in the area of target profiling may be to utilize machine learning models using clinical as well as real world evidence (RWE) data in addition to all available preclinical data for better target and disease validation.
4.ADMET Prediction and Scoring
-
Various academic groups and industry have invested a lot of resources to provide these models due to the fact that there are frequent late stage failures due to either undesirable ADME properties or toxicity issues. Some of these properties could be measured in a high throughput fashion and thereby leading to generation of large data sets suitable for machine learning.
-
It’s imperative to discuss a few best practices:
- models should be interpretable
- models should not only be predictable but provide “confidence” for every prediction
- models should be updated routinely to keep them up to data with newly measured data
- Some sort of prospective predictions should be captured at the time of model update process so that project teams can assess the quality of a model for their projects in a prospective way.
-
An interesting idea to work on would be to build machine learning models that can utilize predicted ADMET properties in addition to physchem properties and generate low dose compounds.
5.Synthesis Planning
- In a more recent work by Coley et al., a panel of ~140K reaction templates was developed as a framework.
- There are several limitations:
- sufficiently cover the reaction space
- insufficient negative examples
- To enable collection of a larger dataset that could potentially contain more diverse and both positive and negative examples, one could imagine building a consortium where various pharmaceutical industry representatives can encrypt their respective ELN datasets and share that publicly at a precompetitive level.
6.Conclusion
- We strongly believe that this is the high time when industry embraces these methods and make them part of their routine drug discovery process.


