热门标签
热门文章
- 1创意十足的网页设计方案案例欣赏_网页设计有创意的主题
- 2利用PP-YOLOE实现雾天环境下的行人车辆目标检测:技术、实现与优化_雾天目标检测
- 3前端加载高德离线地图的解决方案_amaploader实现离线地图
- 4常见开源协议横向对比_开源协议对比
- 5python requests 的代理错误排查:HTTPConnectionPool(host=‘127.0.0.1‘, port=1084)_requests.exceptions.connectionerror: httpconnectio
- 6k8s中文件描述符与线程限制_k8s ulimit
- 7简单HTML网页制作 实例_html网页制作案例
- 8Python中的Base64编码和解码_b64decode
- 9自学 Java 怎么入门?
- 10Xshell7免费版安装教程_xshall7安装教程
当前位置: article > 正文
机器学习-时间序列的特征工程_时间序列输入矩阵有冗余
作者:盐析白兔 | 2024-02-18 07:58:16
赞
踩
时间序列输入矩阵有冗余
In [1]:
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns # more plots
from dateutil.relativedelta import relativedelta # working with dates with style
from scipy.optimize import minimize # for function minimization
import lightgbm as lgb
import xgboost as xgb
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import StandardScaler
from sklearn.preprocessing import MinMaxScaler
from sklearn.model_selection import TimeSeriesSplit
from sklearn.linear_model import LinearRegression
from sklearn.model_selection import cross_val_score
from sklearn.linear_model import Ridge,Lasso
from hyperopt import Trials, STATUS_OK, tpe, hp, fmin
# 忽略warnings
import warnings
warnings.filterwarnings("ignore")
In [2]:
def mean_absolute_percentage_error(y_true, y_pred):
return np.mean(np.abs((y_true - y_pred) / y_true)) * 100
def plotModelResults(model, X_train, X_test, y_train, y_test, plot_intervals=False, plot_anomalies=False, subtitle=""):
"""
Plots modelled vs fact values, prediction intervals and anomalies
"""
prediction = model.predict(X_test)
plt.figure(figsize=(15, 7))
plt.plot(y_test.index, prediction, "g", label="prediction", linewidth=2.0)
plt.plot(y_test.index, y_test.values, label="actual", linewidth=2.0)
if plot_intervals:
tscv = TimeSeriesSplit(n_splits=3)
# for train, test in tscv.split(y_train):
# model.fit(X_train[train],y_train[train])
# B = pd.DataFrame(y_train[test])
# B['pred'] = model.predict(X_train[test])
# score = model.score(X_train[test],y_train[test])
# print(score)
cv = cross_val_score(model, X_train, y_train,
cv=tscv,
scoring="neg_mean_absolute_error")
mae = cv.mean() * (-1)
deviation = cv.std()
scale = 1.96
lower = prediction - (mae + scale * deviation)
upper = prediction + (mae + scale * deviation)
plt.plot(y_test.index, lower, "r--", label="upper bond / lower bond", alpha=0.5)
plt.plot(y_test.index, upper, "r--", alpha=0.5)
if plot_anomalies:
anomalies = np.array([np.NaN] * len(y_test))
anomalies[y_test < lower] = y_test[y_test < lower]
anomalies[y_test > upper] = y_test[y_test > upper]
plt.plot(y_test.index, anomalies, "o", markersize=10, label="Anomalies")
error = mean_absolute_percentage_error(prediction, y_test)
plt.title((subtitle+"Mean absolute percentage error {0:.2f}%").format(error))
plt.legend(loc="best")
plt.tight_layout()
plt.grid(True)
plt.show()
def plotModelBoostingResults(model1, model2, X_test, y_test):
"""
Plots modelled vs fact values, prediction intervals and anomalies
"""
prediction1 = model1.predict(X_test)
prediction2 = model2.predict(X_test)
prediction = prediction1 + prediction2
plt.figure(figsize=(15, 7))
plt.plot(y_test.index, prediction, "g", label="prediction", linewidth=2.0)
plt.plot(y_test.index, y_test.values, label="actual", linewidth=2.0)
error = mean_absolute_percentage_error(prediction, y_test)
plt.title("Mean absolute percentage error {0:.2f}%".format(error))
plt.legend(loc="best")
plt.tight_layout()
plt.grid(True)
plt.show()
def plotCoefficients(model):
"""
Plots sorted coefficient values of the model
"""
coefs = pd.DataFrame(model.coef_, X_train.columns)
coefs.columns = ["coef"]
coefs["abs"] = coefs.coef.apply(np.abs)
coefs = coefs.sort_values(by="abs", ascending=False).drop(["abs"], axis=1)
plt.figure(figsize=(15, 7))
coefs.coef.plot(kind='bar')
plt.grid(True, axis='y')
plt.hlines(y=0, xmin=0, xmax=len(coefs), linestyles='dashed');
def code_mean(data, cat_feature, real_feature):
"""
Returns a dictionary where keys are unique categories of the cat_feature,
and values are means over real_feature
"""
return dict(data.groupby(cat_feature)[real_feature].mean())
def makeCirclyStats(data, target, cols, stats):
for col in cols:
for stat in stats:
data[col + '_' + stat] = data.groupby(col)[target].transform(stat)
def featuresRank(model, X_train):
coefs = pd.DataFrame(model.coef_, X_train.columns)
coefs.columns = ["coef"]
coefs["abs"] = coefs.coef.apply(np.abs)
coefs = coefs.sort_values(by="abs", ascending=False).drop(["abs"], axis=1)
return coefs
def prepareData(data):
data_ = data.copy()
y = data_.dropna().y
X = data_.dropna().drop(['y'], axis=1)
X_train, X_test, y_train, y_test = train_test_split(X, y, shuffle=False, test_size=0.1)
scaler = StandardScaler()
X_train = pd.DataFrame(scaler.fit_transform(X_train),columns=X_train.columns)
X_test = pd.DataFrame(scaler.transform(X_test),columns=X_test.columns)
return X_train, X_test, y_train, y_test
In [3]:
Df = pd.read_excel('/kaggle/input/coal-consume/data.xlsx', index_col=0, date_parser='date')
Df = Df.dropna()
Df.columns = ["y"]
Df.head()
Out[3]:
| y | |
|---|---|
| 指标名称 | |
| 2013-01-01 | 68.6 |
| 2013-01-02 | 68.2 |
| 2013-01-03 | 67.2 |
| 2013-01-04 | 69.4 |
| 2013-01-05 | 71.8 |
In [4]:
data = Df['2015-01-01':].copy()
lags = range(1, 31)
for i in lags:
data["lag_{}".format(i)] = data.y.shift(i)
data.head()
Out[4]:
| y | lag_1 | lag_2 | lag_3 | lag_4 | lag_5 | lag_6 | lag_7 | lag_8 | lag_9 | ... | lag_21 | lag_22 | lag_23 | lag_24 | lag_25 | lag_26 | lag_27 | lag_28 | lag_29 | lag_30 | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 指标名称 | |||||||||||||||||||||
| 2015-01-02 | 66.7 | NaN | NaN | NaN | NaN | NaN | NaN | NaN | NaN | NaN | ... | NaN | NaN | NaN | NaN | NaN | NaN | NaN | NaN | NaN | NaN |
| 2015-01-04 | 68.9 | 66.7 | NaN | NaN | NaN | NaN | NaN | NaN | NaN | NaN | ... | NaN | NaN | NaN | NaN | NaN | NaN | NaN | NaN | NaN | NaN |
| 2015-01-05 | 69.3 | 68.9 | 66.7 | NaN | NaN | NaN | NaN | NaN | NaN | NaN | ... | NaN | NaN | NaN | NaN | NaN | NaN | NaN | NaN | NaN | NaN |
| 2015-01-06 | 72.3 | 69.3 | 68.9 | 66.7 | NaN | NaN | NaN | NaN | NaN | NaN | ... | NaN | NaN | NaN | NaN | NaN | NaN | NaN | NaN | NaN | NaN |
| 2015-01-07 | 72.7 | 72.3 | 69.3 | 68.9 | 66.7 | NaN | NaN | NaN | NaN | NaN | ... | NaN | NaN | NaN | NaN | NaN | NaN | NaN | NaN | NaN | NaN |
5 rows × 31 columns
In [5]:
# 划分训练集测试集
X_train, X_test, y_train, y_test = prepareData(data)
# 线性回归
lr = LinearRegression()
lr.fit(X_train, y_train)
# 画图
plotModelResults(lr, X_train, X_test, y_train, y_test, plot_intervals=True, plot_anomalies=True)
plotCoefficients(lr)

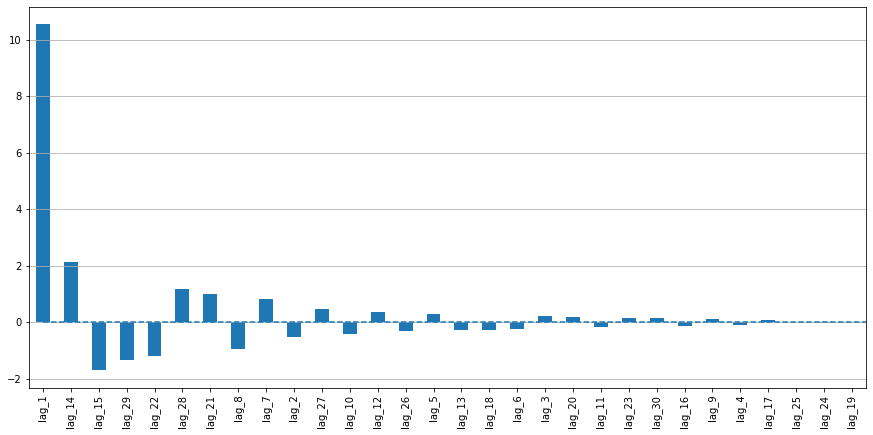
In [6]:
plt.figure(figsize=(10, 8))
sns.heatmap(X_train.corr())
Out[6]:
<AxesSubplot:>

In [7]:
data = Df['2015-01-01':].copy()
lags = [1,14,21,28]
for i in lags:
data["lag_{}".format(i)] = data.y.shift(i)
data.head()
Out[7]:
| y | lag_1 | lag_14 | lag_21 | lag_28 | |
|---|---|---|---|---|---|
| 指标名称 | |||||
| 2015-01-02 | 66.7 | NaN | NaN | NaN | NaN |
| 2015-01-04 | 68.9 | 66.7 | NaN | NaN | NaN |
| 2015-01-05 | 69.3 | 68.9 | NaN | NaN | NaN |
| 2015-01-06 | 72.3 | 69.3 | NaN | NaN | NaN |
| 2015-01-07 | 72.7 | 72.3 | NaN | NaN | NaN |
In [8]:
# 划分训练集测试集
X_train, X_test, y_train, y_test = prepareData(data)
# 线性回归
lr = LinearRegression()
lr.fit(X_train, y_train)
# 画图
plotModelResults(lr, X_train, X_test, y_train, y_test, plot_intervals=True, plot_anomalies=True)
plotCoefficients(lr)


In [9]:
data.index = pd.to_datetime(data.index)
data["weekday"] = data.index.weekday
data['is_weekend'] = data.weekday.isin([5, 6]) * 1
data['quarter'] = data.index.quarter
data['month'] = data.index.month
data['dayofmonth'] = data.index.day
data.head()
Out[9]:
| y | lag_1 | lag_14 | lag_21 | lag_28 | weekday | is_weekend | quarter | month | dayofmonth | |
|---|---|---|---|---|---|---|---|---|---|---|
| 指标名称 | ||||||||||
| 2015-01-02 | 66.7 | NaN | NaN | NaN | NaN | 4 | 0 | 1 | 1 | 2 |
| 2015-01-04 | 68.9 | 66.7 | NaN | NaN | NaN | 6 | 1 | 1 | 1 | 4 |
| 2015-01-05 | 69.3 | 68.9 | NaN | NaN | NaN | 0 | 0 | 1 | 1 | 5 |
| 2015-01-06 | 72.3 | 69.3 | NaN | NaN | NaN | 1 | 0 | 1 | 1 | 6 |
| 2015-01-07 | 72.7 | 72.3 | NaN | NaN | NaN | 2 | 0 | 1 | 1 | 7 |
In [10]:
weekday = code_mean(data, 'weekday', "y")
plt.figure(figsize=(7, 5))
plt.title("weekday averages")
pd.DataFrame.from_dict(weekday, orient='index')[0].plot()
quarter = code_mean(data, 'quarter', "y")
plt.figure(figsize=(7, 5))
plt.title("quarter averages")
pd.DataFrame.from_dict(quarter, orient='index')[0].plot()
month = code_mean(data, 'month', "y")
plt.figure(figsize=(7, 5))
plt.title("month averages")
pd.DataFrame.from_dict(month, orient='index')[0].plot()
dayofmonth = code_mean(data, 'dayofmonth', "y")
plt.figure(figsize=(7, 5))
plt.title("dayofmonth averages")
pd.DataFrame.from_dict(dayofmonth, orient='index')[0].plot()
Out[10]:
<AxesSubplot:title={'center':'dayofmonth averages'}>



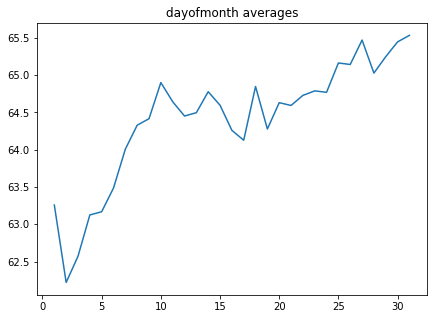
In [11]:
# 划分训练集测试集
X_train, X_test, y_train, y_test = prepareData(data)
# 线性回归
lr = LinearRegression()
lr.fit(X_train, y_train)
# 画图
plotModelResults(lr, X_train, X_test, y_train, y_test, plot_intervals=True, plot_anomalies=True)
plotCoefficients(lr)

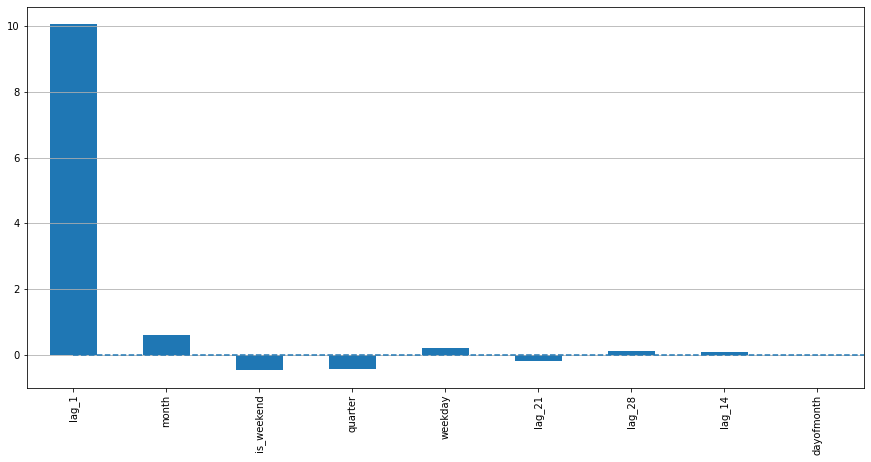
In [12]:
cols = ["weekday", "month", "dayofmonth", "quarter", "is_weekend"]
stats = ["mean","max", "min", "median", "sum", "skew", "std"]
makeCirclyStats(data, "y", cols, stats)
data.head()
Out[12]:
| y | lag_1 | lag_14 | lag_21 | lag_28 | weekday | is_weekend | quarter | month | dayofmonth | ... | quarter_sum | quarter_skew | quarter_std | is_weekend_mean | is_weekend_max | is_weekend_min | is_weekend_median | is_weekend_sum | is_weekend_skew | is_weekend_std | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 指标名称 | |||||||||||||||||||||
| 2015-01-02 | 66.7 | NaN | NaN | NaN | NaN | 4 | 0 | 1 | 1 | 2 | ... | 40348.07 | -0.377789 | 13.283873 | 64.267847 | 93.19 | 29.0 | 63.85 | 108355.59 | -0.046244 | 11.109158 |
| 2015-01-04 | 68.9 | 66.7 | NaN | NaN | NaN | 6 | 1 | 1 | 1 | 4 | ... | 40348.07 | -0.377789 | 13.283873 | 64.644874 | 92.72 | 28.0 | 64.85 | 46027.15 | -0.138350 | 11.039269 |
| 2015-01-05 | 69.3 | 68.9 | NaN | NaN | NaN | 0 | 0 | 1 | 1 | 5 | ... | 40348.07 | -0.377789 | 13.283873 | 64.267847 | 93.19 | 29.0 | 63.85 | 108355.59 | -0.046244 | 11.109158 |
| 2015-01-06 | 72.3 | 69.3 | NaN | NaN | NaN | 1 | 0 | 1 | 1 | 6 | ... | 40348.07 | -0.377789 | 13.283873 | 64.267847 | 93.19 | 29.0 | 63.85 | 108355.59 | -0.046244 | 11.109158 |
| 2015-01-07 | 72.7 | 72.3 | NaN | NaN | NaN | 2 | 0 | 1 | 1 | 7 | ... | 40348.07 | -0.377789 | 13.283873 | 64.267847 | 93.19 | 29.0 | 63.85 | 108355.59 | -0.046244 | 11.109158 |
5 rows × 45 columns
In [13]:
# 划分训练集测试集
X_train, X_test, y_train, y_test = prepareData(data)
# 线性回归
lr = LinearRegression()
lr.fit(X_train, y_train)
# 画图
plotModelResults(lr, X_train, X_test, y_train, y_test, plot_intervals=True, plot_anomalies=True)
plotCoefficients(lr)

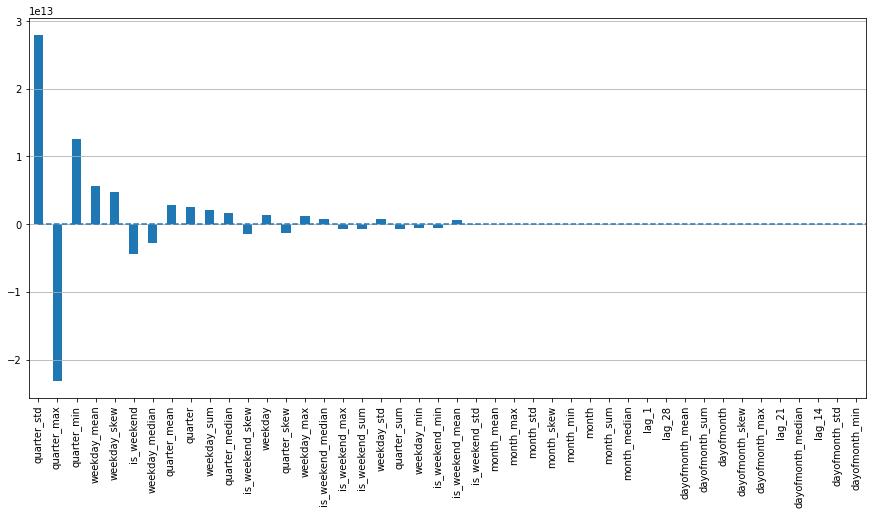
In [14]:
data.columns
Out[14]:
Index(['y', 'lag_1', 'lag_14', 'lag_21', 'lag_28', 'weekday', 'is_weekend',
'quarter', 'month', 'dayofmonth', 'weekday_mean', 'weekday_max',
'weekday_min', 'weekday_median', 'weekday_sum', 'weekday_skew',
'weekday_std', 'month_mean', 'month_max', 'month_min', 'month_median',
'month_sum', 'month_skew', 'month_std', 'dayofmonth_mean',
'dayofmonth_max', 'dayofmonth_min', 'dayofmonth_median',
'dayofmonth_sum', 'dayofmonth_skew', 'dayofmonth_std', 'quarter_mean',
'quarter_max', 'quarter_min', 'quarter_median', 'quarter_sum',
'quarter_skew', 'quarter_std', 'is_weekend_mean', 'is_weekend_max',
'is_weekend_min', 'is_weekend_median', 'is_weekend_sum',
'is_weekend_skew', 'is_weekend_std'],
dtype='object')
In [15]:
data = data.drop(columns=['weekday_mean', 'month_mean',
'dayofmonth_mean', 'quarter_mean', 'is_weekend_mean', 'weekday_max',
'weekday_min', 'weekday_median', 'weekday_sum', 'weekday_skew',
'weekday_std', 'month_max', 'month_min', 'month_median', 'month_sum',
'month_skew', 'month_std', 'dayofmonth_max', 'dayofmonth_min',
'dayofmonth_median', 'dayofmonth_sum', 'dayofmonth_skew',
'dayofmonth_std', 'quarter_max', 'quarter_min', 'quarter_median',
'quarter_sum', 'quarter_skew', 'quarter_std', 'is_weekend_max',
'is_weekend_min', 'is_weekend_median', 'is_weekend_sum',
'is_weekend_skew', 'is_weekend_std'])
In [16]:
data['rolling_mean_7'] = data.rolling(7)['y'].mean()
data['rolling_mean_14'] = data.rolling(14)['y'].mean()
data['rolling_mean_28'] = data.rolling(28)['y'].mean()
In [17]:
# 划分训练集测试集
X_train, X_test, y_train, y_test = prepareData(data)
# 线性回归
lr = LinearRegression()
lr.fit(X_train, y_train)
# 画图
plotModelResults(lr, X_train, X_test, y_train, y_test, plot_intervals=True, plot_anomalies=True)
plotCoefficients(lr)
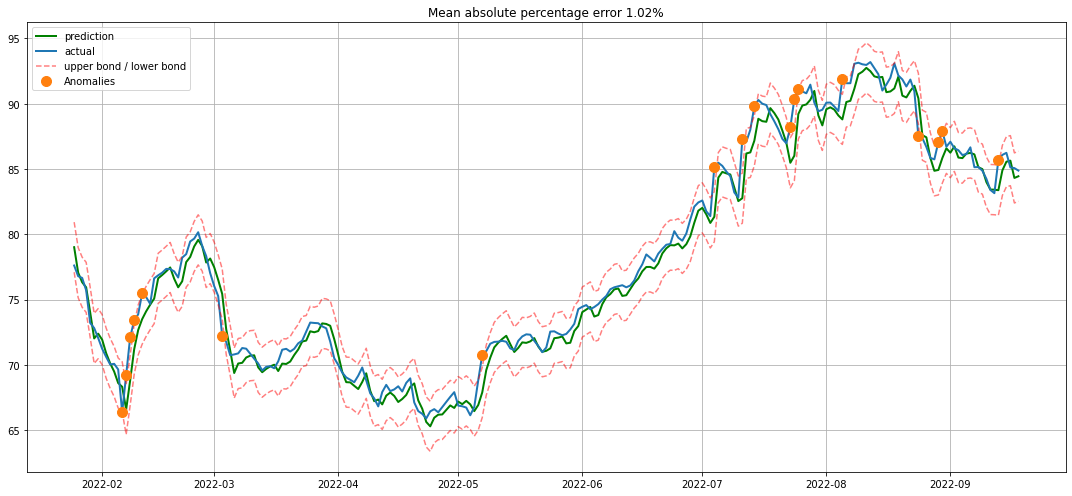
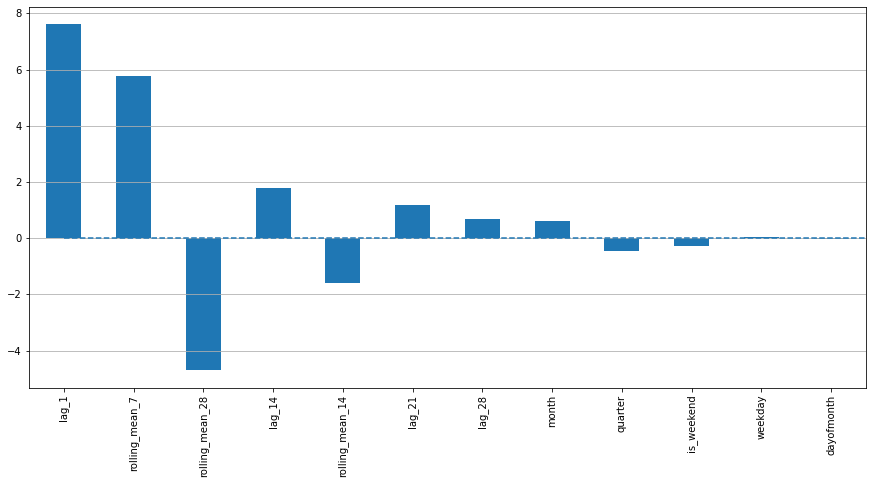
In [18]:
data['week_rolling_mean_4'] = data.groupby('weekday')["y"].transform(lambda x: x.rolling(4).mean())
In [19]:
# 划分训练集测试集
X_train, X_test, y_train, y_test = prepareData(data)
# 线性回归
lr = LinearRegression()
lr.fit(X_train, y_train)
# 画图
plotModelResults(lr, X_train, X_test, y_train, y_test, plot_intervals=True, plot_anomalies=True)
plotCoefficients(lr)

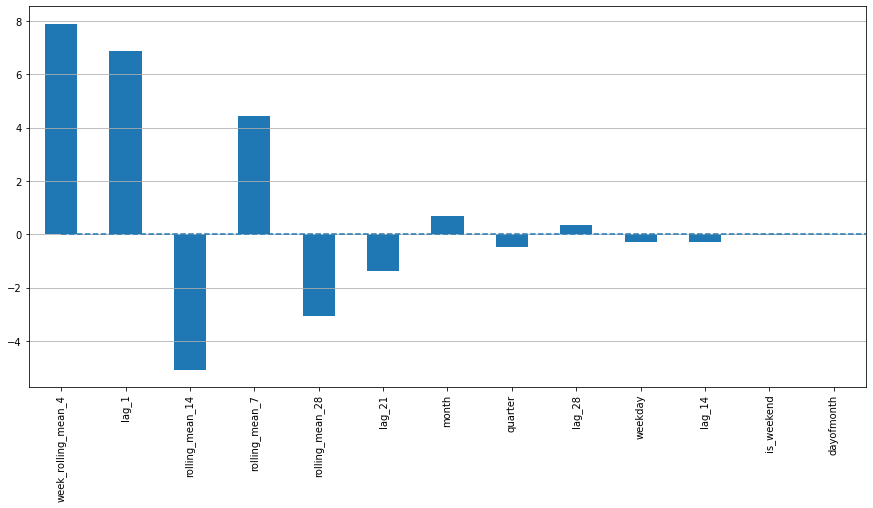
In [20]:
data['month_expanding_mean'] = data.groupby('month')["y"].transform(lambda x: x.expanding(10).mean())
In [21]:
# 划分训练集测试集
X_train, X_test, y_train, y_test = prepareData(data)
# 线性回归
lr = LinearRegression()
lr.fit(X_train, y_train)
# 画图
plotModelResults(lr, X_train, X_test, y_train, y_test, plot_intervals=True, plot_anomalies=True)
plotCoefficients(lr)


In [22]:
data = data.drop(columns=['month_expanding_mean','dayofmonth','is_weekend'])
In [23]:
# 划分训练集测试集
X_train, X_test, y_train, y_test = prepareData(data)
# 线性回归
lr = LinearRegression()
lr.fit(X_train, y_train)
# 画图
plotModelResults(lr, X_train, X_test, y_train, y_test, plot_intervals=True, plot_anomalies=True)
plotCoefficients(lr)


In [24]:
plt.figure(figsize=(10, 8))
sns.heatmap(X_train.corr())
Out[24]:
<AxesSubplot:>

In [25]:
featuresRank(lr, X_train)
Out[25]:
| coef | |
|---|---|
| week_rolling_mean_4 | 7.910972 |
| lag_1 | 6.853260 |
| rolling_mean_14 | -5.050413 |
| rolling_mean_7 | 4.433046 |
| rolling_mean_28 | -3.149058 |
| lag_21 | -1.368408 |
| month | 0.692110 |
| quarter | -0.496375 |
| lag_28 | 0.370576 |
| weekday | -0.322907 |
| lag_14 | -0.264314 |
In [26]:
lr.intercept_
Out[26]:
62.794430379746814
In [27]:
y_train_e = y_train - lr.predict(X_train)
y_train_e.head()
Out[27]:
指标名称
2015-01-31 -0.142392
2015-02-01 -0.748898
2015-02-02 1.780891
2015-02-03 -0.680682
2015-02-04 -1.622106
Name: y, dtype: float64
In [28]:
from sklearn.model_selection import cross_val_score
def objective(space):
model = xgb.XGBRegressor(n_estimators=space['n_estimators'],
max_depth=int(space['max_depth']),
learning_rate=space['learning_rate'],
gamma=space['gamma'],
min_child_weight=space['min_child_weight'],
subsample=space['subsample'],
colsample_bytree=space['colsample_bytree']
)
model.fit(X_train, y_train_e)
# Applying k-Fold Cross Validation
tscv = TimeSeriesSplit(n_splits=3)
score = cross_val_score(estimator=model, X=X_train, y=y_train_e, cv=tscv)
CrossValMean = score.mean()
return {'loss': 1 - CrossValMean, 'status': STATUS_OK}
space = {
'max_depth': hp.choice('max_depth', range(5, 30, 1)),
'learning_rate': hp.quniform('learning_rate', 0.01, 0.5, 0.01),
'n_estimators': hp.choice('n_estimators', range(20, 205, 5)),
'gamma': hp.quniform('gamma', 0, 0.50, 0.01),
'min_child_weight': hp.quniform('min_child_weight', 1, 10, 1),
'subsample': hp.quniform('subsample', 0.1, 1, 0.01),
'colsample_bytree': hp.quniform('colsample_bytree', 0.1, 1.0, 0.01)}
In [29]:
# trials = Trials()
# best = fmin(fn=objective,
# space=space,
# algo=tpe.suggest,
# max_evals=50,
# trials=trials)
In [30]:
# print(best)
In [31]:
# xgbReg = xgb.XGBRegressor(n_estimators=best['n_estimators'],
# max_depth=best['max_depth'],
# learning_rate=best['learning_rate'],
# gamma=best['gamma'],
# min_child_weight=best['min_child_weight'],
# subsample=best['subsample'],
# )
xgbReg = xgb.XGBRegressor(n_estimators=20,
max_depth=22,
learning_rate=0.01,
gamma=0.19,
min_child_weight=7,
subsample=0.65,
colsample_bytree=0.14
)
xgbReg.fit(X_train, y_train_e)
Out[31]:
XGBRegressor(base_score=0.5, booster='gbtree', callbacks=None,
colsample_bylevel=1, colsample_bynode=1, colsample_bytree=0.14,
early_stopping_rounds=None, enable_categorical=False,
eval_metric=None, gamma=0.19, gpu_id=-1, grow_policy='depthwise',
importance_type=None, interaction_constraints='',
learning_rate=0.01, max_bin=256, max_cat_to_onehot=4,
max_delta_step=0, max_depth=22, max_leaves=0, min_child_weight=7,
missing=nan, monotone_constraints='()', n_estimators=20, n_jobs=0,
num_parallel_tree=1, predictor='auto', random_state=0, reg_alpha=0,
reg_lambda=1, ...)
In [32]:
plotModelBoostingResults(lr, xgbReg, X_test, y_test)
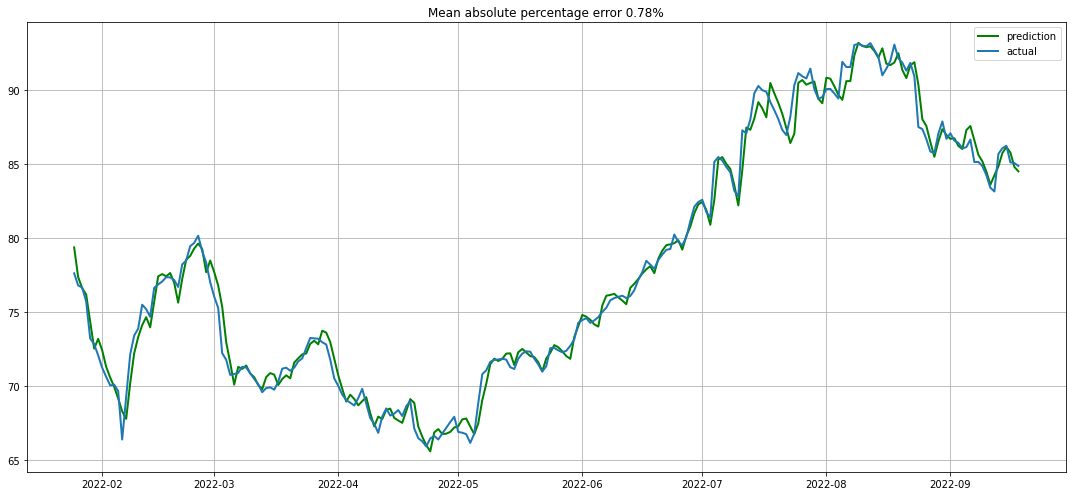
声明:本文内容由网友自发贡献,不代表【wpsshop博客】立场,版权归原作者所有,本站不承担相应法律责任。如您发现有侵权的内容,请联系我们。转载请注明出处:https://www.wpsshop.cn/w/盐析白兔/article/detail/106309
推荐阅读
相关标签



